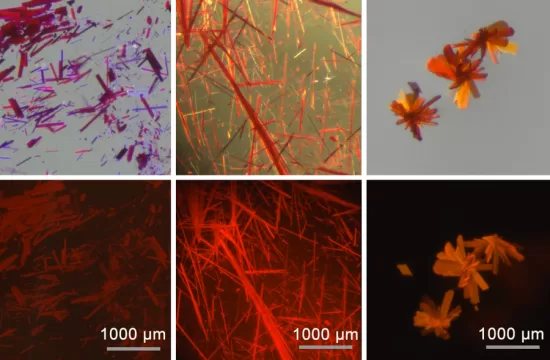
Taipei, March 23 – Carbohydrates are among the most complex biomolecules in nature, but analyzing their structures has long posed challenges for scientists. A team at National Taiwan University has now unveiled a new analytical platform that could make the process faster and more precise.
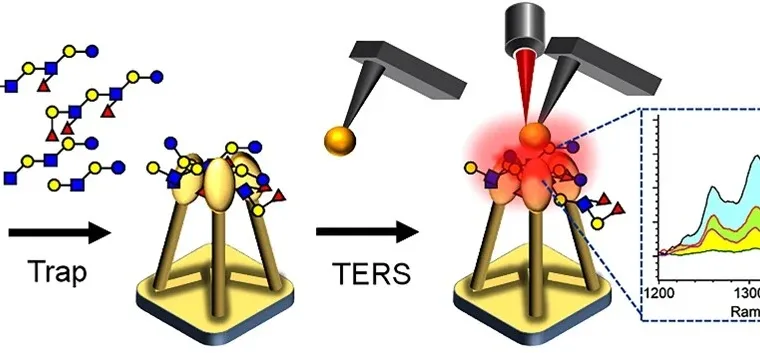
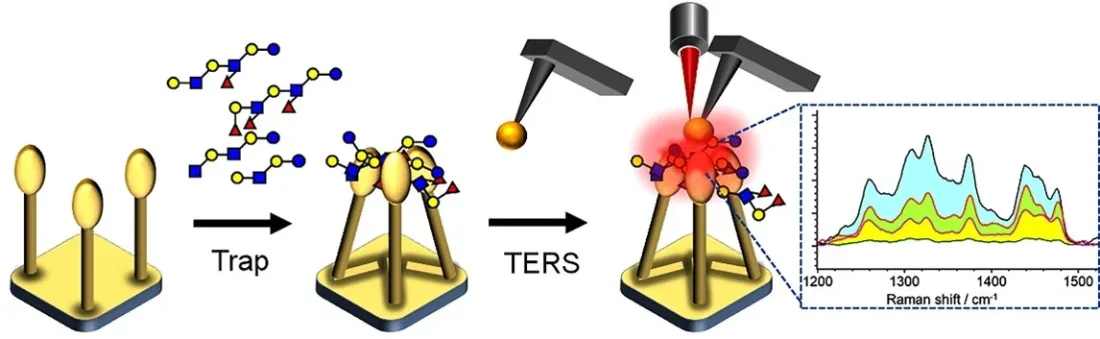
The method, published in Angewandte Chemie International Edition, combines tip‑enhanced Raman spectroscopy (TERS) with nanopillar‑assisted glycan trapping. By confining glycans in a localized plasmonic region, the system produces strong vibrational signals from minute samples, enabling label‑free detection of structural details.
Researchers said the platform can distinguish subtle differences in glycosidic linkages, such as β(1→3) and β(1→4), which are difficult to separate using conventional techniques. The team also reported that characteristic H–C–H deformation signals can help estimate chain length, while an azido tag improves signal normalization when only trace volumes are available.
Beyond static analysis, the system can monitor biosynthesis in real time. The study demonstrated enzymatic glycosylation reactions, including the appearance of α1→2 fucosylation linkages during product formation.
“This work shows that tip‑enhanced Raman spectroscopy can move beyond simple detection and become a practical tool for decoding carbohydrate structures with very high specificity,” said Dr. Kien Voon Kong, co‑corresponding author and professor of chemistry at National Taiwan University.
The researchers said the platform could open new opportunities in glycoscience, biomaterials and diagnostics, where understanding both molecular identity and biosynthetic progression is critical.